A technology called machine learning is behind that seemingly magical ability of social networking websites to identify people in posted photos. By analyzing subtle patterns in facial features, machine learning algorithms can recognize people you’ve never tagged and might not even know.
Machine learning is also transforming how scientists at Novartis discover and develop new drugs. Similar to how social networking websites use the technology to classify people on your computer screen, Novartis scientists use it to classify digital images of cells, each treated with different experimental compounds. The algorithms group compounds with similar effects visually and with lightning speed. Biological insights that might take months to generate using time-consuming laboratory experiments and human visual inspection can be revealed much faster using automated computer algorithms looking at pictures. As a result, machine learning has the potential to shrink drug discovery timelines, which could help patients get quicker access to new therapies.
Machine learning has the potential to shrink drug discovery timelines, which could help patients get quicker access to new therapies.
“Machine learning is pointing us to new therapeutic possibilities with unprecedented efficiency,” says Jeremy Jenkins, Head of Informatics for Chemical Biology and Therapeutics at the Novartis Institutes for BioMedical Research (NIBR). “And it has an unparalleled ability to teach us about how our drugs are working.”
Novartis scientists recently pioneered advances in a specific type of machine learning called deep learning, which mimics how the eyes and brain process visual images. Our eyes perceive light in different shades, and tightly coordinated neuronal networks transform those patterns into the colors and shapes that we associate with familiar objects, faces, and other living things in our surroundings. Taking a cue from nature, the Novartis team simulated that same approach and taught a computerized neuronal network how to recognize the subtle changes that experimental compounds induce in a cell.
The team initially used a supervised approach to deep learning, meaning they had to teach the system how to recognize particular effects from treatment – such as changes in a cell’s shape or in the activity of its proteins – before the system could recognize those effects on its own. They trained the network by showing it images of cells that were treated with compounds known to work in a particular way, so that it would learn the visual patterns associated with the different drug mechanisms. Then they tested it using pictures of cells treated with over 100 mystery compounds.
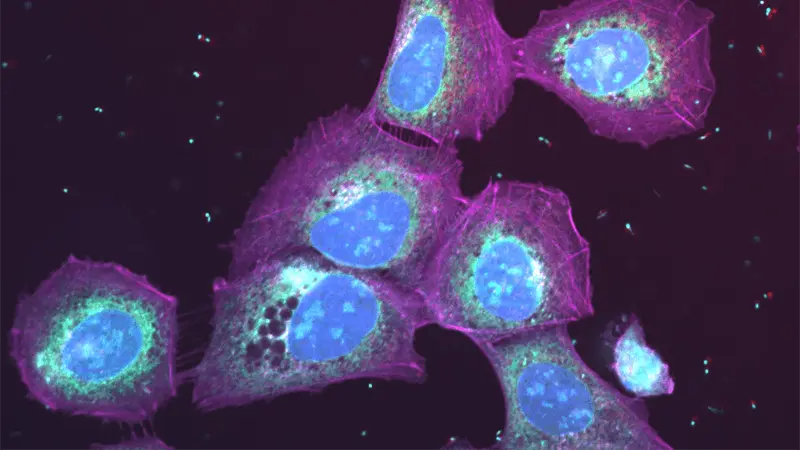
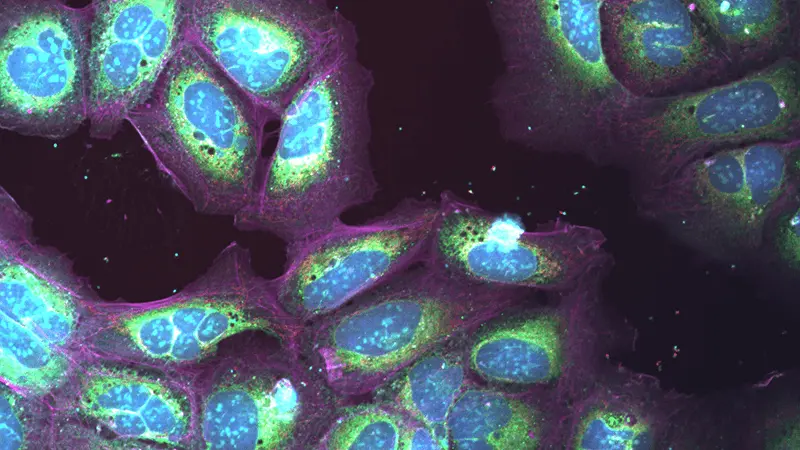
The computerized network predicted correctly how the chemicals would affect the cells, even at different doses. “That shows it’s possible for the system to go from a digital picture to a biological understanding of what the drugs are doing in a single step,” says William Godinez, a lab head at NIBR’s Infectious Diseases Department in Emeryville, California, who led the research. “The predictions were nearly 100% accurate.” The Novartis researchers published their approach and results in the journal Bioinformatics last year.
The predictions were nearly 100% accurate.
More recently, Godinez and others at Novartis have made important strides with unsupervised systems that don’t require any initial instructions. Unsupervised algorithms sort through and classify pictures of treated cells automatically.
They also reveal biological changes – some with clinical potential – that scientists might not previously have thought to look for. “Since we’ve historically looked for specific types of effects, we’ve been held back by the limits of our prior knowledge,” explains Xian Zhang, an investigator in Jenkins’ group at NIBR who was also Godinez’ postdoc advisor.
Unsupervised machine learning systems remove those limits because they aren’t bound by prior assumptions about how different compounds affect cells. They simply sort the images and group them by shared visual patterns. “The algorithms don’t know what they’re looking at, but that doesn’t matter,” Zhang says. “They find the differences between cells that allow us to form new hypotheses to test.”
Opportunities for machine learning extend from early-stage drug discovery through testing in patients during clinical trials, Jenkins says.
One time-consuming part of drug discovery is testing compounds against samples of diseased cells – a process that often requires painstaking analyses of each sample to find compounds that are biologically active and worth further investigation. To speed this screening process, the team is using images from these time-intensive experiments to train machine learning algorithms to rapidly predict which untested compounds might be worth more study. They are starting with a collection of 3000 compounds but aim to eventually expand the use of machine learning to screen all the approximately 1.5 million compounds in the Novartis archive. “We need to do smarter screening, not just larger screening,” says Jenkins.
Unsupervised machine learning systems remove those limits because they aren’t bound by prior assumptions about how different compounds affect cells.
Machine learning algorithms could also be used to sort through various types of medical images during clinical trials and match features of this data to patient responses to the therapy. The algorithms might then have the potential to predict how future patients will respond to the experimental treatment, giving the researchers information that could help them focus testing of the compound on patients who will benefit most.
Pharmaceutical research is complex, and end-to-end drug discovery via computer simulation won’t occur overnight, nor will a single company accomplish it. “We still have a lot to learn about how machine learning can be applied under different scenarios and in different contexts,” cautions Zhang.
Still, says Jenkins, “Machine learning is poised to accelerate a number of critical steps, and we think it could speed up discovery and development for many of our projects.”
Main image: Courtesy Shutterstock/spainter_vfx.
Learn how Novartis is using #machinelearning to accelerate #drugdiscovery.