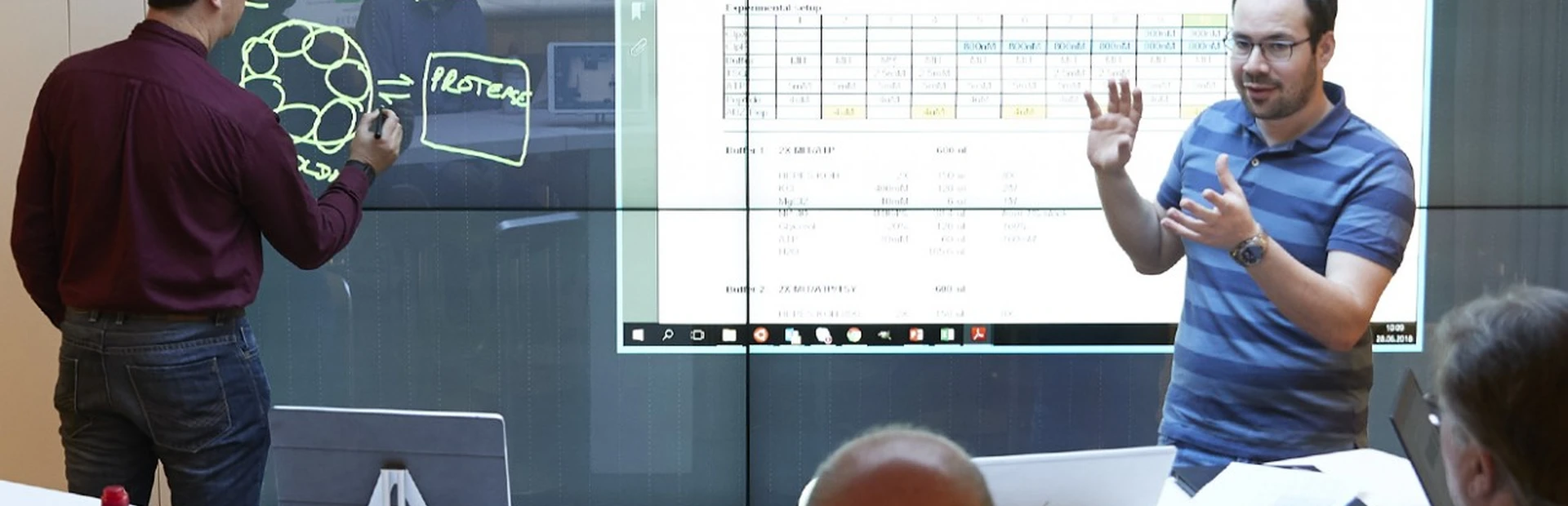
“I vividly remember that conversation as one of the most extraordinary brainstorming sessions I have ever had,” says Hans Voshol, a senior investigator and protein expert who met with Edward Oakeley at a Christmas party back in 2015. “One of the kinds where you can come up with ideas no matter how unlikely they are. Textbook innovative thinking, I guess.”
The two Novartis scientists shared a common interest in proteins – or, more so, a real passion for proteins. Voshol had studied biology in the Netherlands and had specialized in protein biochemistry during his studies. He joined Novartis in 1996 and has worked in this area ever since. Oakeley, an Anglo-Swiss biochemist, established the Friedrich Miescher Institute (FMI) functional genomics facility and helped found the Next Generation Sequencing groups in both Biomarker Development and Analytical Sciences & Imaging (AS&I) at Novartis.
Unwinding a ball of string
On that December evening three years ago, the scientists’ lively discussion was all about enzymes and their ability to untangle proteins. “In simple words, you can think of a protein as a ball of string with knots in it. There are pieces that are very highly tangled and other regions that are much less so,” explains Oakeley. “The enzyme I was looking for should be able to unravel this.”
The two discussed how this enzyme should also use a fluorescent fuel. They imagined that, like old Morse code, the pattern and intensity of the light flashes from this fluorescent fuel could be used to identify and quantify which proteins are in a mixture, and perhaps provide insights into their structures.
For these measuring experiments, Oakeley intended to use an analytical instrument that was originally designed by Pacific Biosciences for DNA sequencing. After five years of lab service, this device was reaching the end of its life span and, if no other use for it could be found, was destined for disposal.
Oakeley’s vision was to repurpose this instrument so it could be used to record the unwinding of thousands to millions of protein molecules at a time. These recordings should not only allow for measuring the quantity of each protein but also for identifying their structures – something which is traditionally rather difficult to do.
However, the key question still was whether there are enzymes that could do the unwinding. If one could be found, then perhaps the life of the “old” sequencing machine could be saved. So it came as a great Christmas present to Oakeley when Voshol said, “I think CLPX is exactly what you need. Why don’t you Google it?”
And so he did. CLPX is an enzyme that can unwind any tagged protein and that uses a readily available fuel, adenosine triphosphate (ATP), as an energy source. ATP could also be fluorescently labeled, which sounded ideal. This was just what Oakeley had been looking for. On the same evening, he summarized everything in a mind map, filed it away, and went to work as usual.
From dream to reality
Nothing happened for two years until Novartis announced the start of Genesis Labs, its internal research startup platform. Upon hearing about this, Oakeley immediately teamed up with Mevion Oertli, Ulrike Naumann and Stefan Reinker from AS&I and with Marc Sultan from Biomarker Development to submit the Single Molecule Next Generation Sequencing idea, which later was picked as one of the winning projects to receive funding.
The pitch jury was impressed. Never before had they heard or thought of such an idea. If it could be implemented successfully, it would take protein analysis to a new level, they believed.
Oakeley and his colleagues started to work and quickly identified several key hurdles to reach their goal. “Our project comprises three different challenges,” Oakeley says. “One is the instrument itself; it is an old system at the end of its productive life, and part of the motivation for our project was to find something new to do with it rather than scrapping it.”
Second, the core team also had to organize all of the reagents that are needed to run their experiments, since there are no standard solutions or kits available. Everything had to be custom-synthesized, which required collaboration across the company. Oakeley and Oertli reached out to their colleagues in other functions.
One of the many collaborators, Marica Pettinelli – a member of Microscopy and Biophotonics – supported the project by trying to make exotic versions of ATP, which were tested as measurable fuel molecules. “This project has an ultimate ambitious goal, and the passion coming from the team has always been energizing for me,” she says. “Genesis Labs can really create connections and opportunities that would be otherwise difficult to establish.”
The third challenge centered on creating a new software solution to sift through the data.
Filtering back to the organization
More than a year into the project, the core team gathered a lot of experience and established efficient processes to reach the goal Oakeley had envisaged at the Christmas party.
We had many ups and downs. It may take us 18 months to learn how to perform a five-minute experiment!
“We do experiments every day, sometimes on the sequencing machine, or just biochemistry experiments in the lab,” says Oertli. “We have many ups and downs. It may take us 18 months to learn how to perform a five-minute experiment!”
Because every Genesis project has a time limit, there is some uncertainty about what will happen after the 18-month project period ends. “If the project shows value, which we are convinced of, then we will feed these findings back into the organization. And we have already uncovered many things that were unexpected,” says Oakeley.
Working on this Genesis Labs project always reminds Oakeley of something that Barbara Hohn, a former group leader at the FMI, used to say: “Potential discoveries grow like flowers on the side of the road. Open your eyes or you will miss them as you rush on by.”

In simple words, you can think of a protein as a ball of string with knots in it. There are pieces that are very highly tangled and other regions that are much less so. The enzyme I was looking for should be able to unravel this.
Edward Oakeley
Novartis scientists are reimagining protein sequencing – with an old device.